Legionella PCR Test
EST’s NEW Quantitative Viable Legionella vPCR
The Legionella Testing Revolution has ARRIVED!
Culture Legionella methods can take a week or more to be reported back to the customer. This can cause a critical delay in establishing a remediation protocol and returning a building to full use after a shutdown. Polymerase Chain Reaction (PCR) rapid testing for Legionella has been available for many years, but PCR does not distinguish living from dead Legionella leading to potential false positives that require a culture to confirm, resulting in the same costly delays. PCR tests that only remove “dead” free DNA (via filtration or enzymatic processing of the sample) do NOT remove Legionella DNA from intact dead bacteria in the sample.
When it comes to testing water for Legionella!
Speed can save lives!
Our quantitative viable Legionella PCR Test provides you with LIVING Legionella counts (CFU/ml*) in as little as 6 HOURS! This is living Legionella only (PMA exclusion) , no false positives due to intact dead Legionella or free DNA, no need to confirm with culture. This analysis will provide you the quickest most reliable confirmation sampling for Legionella allowing you to quickly proceed with remediation or reopening of your building.

But wait, there is more!
Why wait a week for your viable Legionella culture results?
Next Day or Same Day Turnaround available
The test is offered with two different turnaround times allowing for additional information to be combined with this analysis.
NEXT DAY results are reported as CFU/ml* for Legionella pneumophila at the limit of detection (LOD) as described above reported on the next business day. There is an option with this analysis to also request a Next Day Total Bacteria Count and/or a Next Day Pseudomonas aeruginosa screening.
SAME DAY results are reported as CFU/ml* for Legionella pneumophila at the limit of detection (LOD) as described above and are reported 6 hours after receipt in the Laboratory.
*Note that the results are given in units of CFU/ml so you can use the numbers just like you do with culture results. Clearly there are no “colonies” to count in a PCR procedure, so no actual Colony -Forming Units (CFU) are counted. But, the Genomic Units (GU) that are detected in the PCR reaction are from DNA that has been extracted and polymerized from viable Legionella only. Thus, in a Legionella vPCR reaction a GU is equivalent to a CFU, and as such is an infectious unit capable of causing Legionaries’ Disease.
Viability PCR with PMA and PMAxxTM dyes

How does it work?
Viability PCR (vPCR) is a powerful technology for the sensitive and rapid detection of viable microorganisms. Unlike time-consuming culturing methods, qPCR is a fast and sensitive method of detection. However, normal PCR does not distinguish between live and dead cells.
With v-PCR using PMA, you get the speed, sensitivity and specificity of qPCR, plus quantified viability. And because no culturing is required, you can even detect viable but not culturable (VBNC) bacteria. The vPCR technology can be applied not only to bacteria but to other organisms like yeast, and parasites.
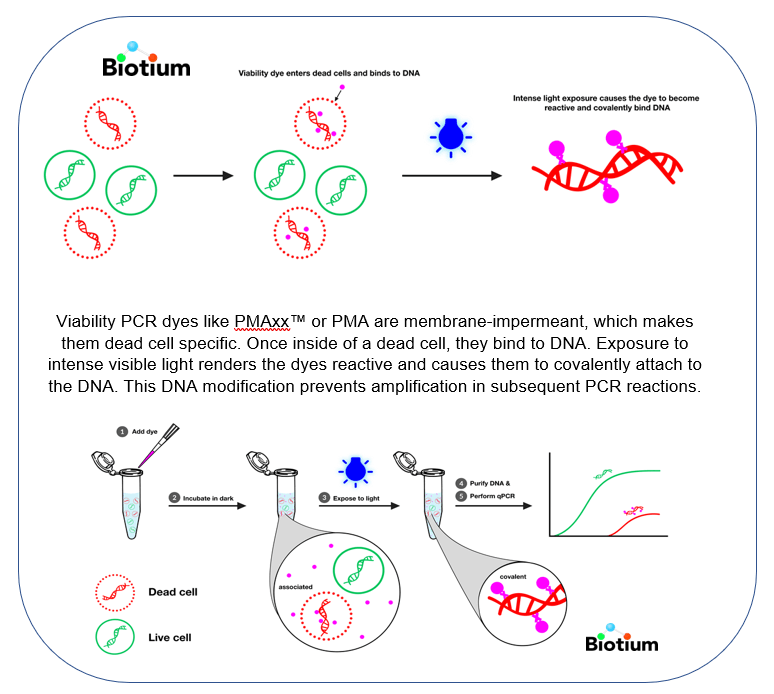
As the inventor of both PMAxx™ and PMA dyes for v-PCR, Biotium offers the expertise, and the widest selection of v-PCR dyes, kits and accessory products.
https://biotium.com/technology/microbiology/pma-for-viability-pcr/
https://biotium.com/wp-content/uploads/2013/07/PMAxx-PMA-Flyer.pdf
https://biotium.com/wp-content/uploads/2015/05/PI-40069.pdf
The Science has Spoken:
Scientific Peer-Reviewed Publications
The science is in! Over 15 years of scientific publications have now confirmed these living versus dead findings with PMA, and with the new and improved PMAxx™, using a variety of bacteria, archaea, yeast, eukaryotic parasites, and even enveloped viruses! Complex matrices, such as mixed microbial biofilms have also been analyzed with the same confirmatory results.
Legionella Viable PCR Publications
Yanez, M.A., A. Nocker, E. Soria-Soria, R. Murtula, L. Martinez and V. Catalan. 2011. Quantification of viable Legionella pneumophila cells using propidium monoazide combined with quantitative PCR. J. Microbiol. Methods. 85: 124-130.
Ditommaso, S, E. Ricciardi, M. Giacomuzzi, S.R. Arauco Rivera, A. Ceccarelli, and C.M. Xotti. 2014. Overestimation of the Legionella spp. load in environmental samples by quantitative real-time PCR: Pretreatment with propidium monoazide as a tool for the assessment of an association between Legionella concentration and sanitary risk. Diagn. Microbiol. Infect. Dis. 80: 260-266.
Li, H. H. Xin, and S.F.Y. Li. 2015. Multiplex PMA-qPCR assay with internal amplification control for simultaneous detection of viable Legionella pneumophila, Salmonella typhimurium, and Staphylococcus aureus in environmental waters. Environ. Sci. Technol. 49: 14249-14256.
Delgado-Viscogliosi, P., L. Solignac, and J.-M. Delattre. 2015. Viability PCR, a culture-independent method for rapid and selective quantification of viable Legionella pneumophila cells in environmental water samples. Appl. Environ. Microbiol. 75: 3502-3512.
Ditommaso, S., M. Giacomuzzi, E. Ricciardi, and C.M. Zotti. 2016. Cultural and molecular evidence of Legionella spp. colonization in dental unit waterlines: Which is the best method for risk assessment? Int. J. Environ. Res. Pub. Health. 13: 211.
Scaturro, M., S. Fontana, I. Dell’eva, et.al. 2016. A multicenter study of viable PCR using propidium monoazide to detect Legionella in water samples. Diagn. Microbiol. Infect. Dis. 85: 283-288.
Bonetta, S., C. Pignata, S. Boneta, et al. 2017. Viability of Legionella pneumophila in water samples: a comparison of propidium monoazide (PMA) treatment on membrane filters and in liquid. Int. J. Environ. Res. Publ. Hlth. 14: 467-473.
Lizana, X, A. Lopez, S. Benito, G. Agusti, M. Rios, N. Pique, and A. M. Marques. 2017. Viability qPCR, a new tool for Legionella risk management. Int J. Hyg. Environ. Hlth. 220: 1318-1324.
Other Significant Viable PCR Publications
Nocker, A., C.-Y. Cheung, and A.K. Camper. 2006. Comparison of propidium monoazide with ethidium monoazide for differentiation of live vs. dead bacteria by selective removal of DNA from dead cells. J. Microbiol. Methods. 67: 310-320.
Nocker, A., P. Sossa-Fernandez, M.D. Burr, and A.K. Camper. 2007. Use of propidium monoazide for live/dead distinction in microbial ecology. Appl. Environ. Microbiol. 73: 5111-5117.
Check out 200 references of this published science on the Biotium website:
https://biotium.com/wp-content/uploads/2017/10/PMA-PMAxx-References.pdf



